Harnessing the power of big data holds promise for equine precision medicine
It might seem a paradox. By starting with an enormous amount of data, data sets so large they can only be explored via supercomputers, there is now the possibility of customized medicine precisely tailored to a single, individual horse.
This innovation in equine medicine follows the work that has been going on in human medicine. It is built on the foundation of the first whole-genomic DNA sequence of the horse, called the Equine Reference Genome, an accomplishment widely considered to be one of the most important in equine science to date.
Behind the computer screens, finding patterns in the hundreds of terabytes of data generated through genetic and genomic studies on the horse, is Ted Kalbfleisch, PhD, associate professor at the University of Kentucky’s Gluck Equine Research Center.
Kalbfleisch specializes in bioinformatics, an interdisciplinary field that combines biology, computer science, information science, mathematics and statistics to analyze and interpret biological data. The growing need for and promise of that interpretation is why Kalbfleisch’s name appears next to the researchers leading several of the studies currently being done at the Gluck Center that have a genetic or genomic component to them. (Genomics is the study of all the DNA, or the genome, including genes and interactions of those genes with each other and the horse’s environment.)
“Those reference genomes are the foundation of nearly all genetic and genomic work that is being done in horses, and in fact, across nearly all human health related organisms and agricultural species. The reference genomes that have emerged in the last decade have been transformative in the way we do science,” he said. “The word unprecedented is thrown around a lot these days, but to be clear, the revolution that assembled genomes have driven in health science is truly without precedent in any other field of science. All that said, these are still very early days in this new era.”
According to Kalbfleisch, the ultimate objective of using “big data” and the promise it holds for equine health outcomes is what is being called “precision medicine” in clinical applications with both human and veterinary patients.
He said he looks at it this way, “Given everything that can be known about an animal, its genome, the genomes of the bacteria working away in the animal’s gut, what can we do specifically for that animal to increase the odds that it lives a healthy, productive life? And if it does become ill, what can we do to make certain the treatment is tailored to that animal to ensure recovery without side effects, or other collateral damage?”
In a chapter he and James MacLeod, VMD, PhD, John S. and Elizabeth A. Knight chair at the Gluck Center and director of UK Ag Equine Programs, co-wrote for a recently published textbook (Vet Clin North Am Equine Pract. 2020 Aug;36(2):173-181. doi: 10.1016/j.cveq.2020.04.002), they explained that the applied aspects of genomics are now coming to fruition in many areas of equine science and veterinary medicine, including the emergence of what they call equine precision P4 medicine: predictive, preventative, personalized and participatory.
An excerpt from that book chapter:
This is the dawn of P4 medicine. Fortunately for horses and the people who care about them, equine science and equine veterinary medicine are well-positioned to participate. Precision medicine, earlier referred to as personalized medicine, is now an established concept that has become a major driver for transformative changes underway in health care and biomedical research. Each individual patient, as opposed to a large group or population of patients, is the focus. This is true for human and veterinary patients, including prophylactic, diagnostic, therapeutic and patient monitoring applications.
Shifting the emphasis of health care from treating disease (reaction) to maintaining health (prevention), or at least balancing these relationships, is a foundation of the changes underway.
The Equine Genome
The first version of the human reference genome was published at the beginning of 2000. This achievement took approximately 10 years, the efforts of hundreds of scientists and a global investment of several billions of dollars to complete.
A horse has 31 pairs of autosomal chromosomes, half coming from the sire and the other half from the dam. Each horse’s genome has about 2.5 billion bases of DNA. The first whole genomic DNA sequence of the horse, a map of the DNA of a Thoroughbred mare named Twilight, was completed in 2007 and published in 2009. This achievement included contributions from MacLeod and Ernest Bailey, PhD, professor at the Gluck Center. In 2014, the equine genomics research community began a project to improve the reference sequence for the horse, a project led by Kalbfleisch that included contributions from MacLeod and Bailey.
By re-analyzing DNA from the original horse reference genome, scientists generated a more than tenfold increase in data and types of data to correct thousands of errors in the original sequence. In the years since the first equine genome was published, costs associated with generating genome sequences have also seen a change. Now, for around $1,000 and the appropriate computer software and expertise, a genome sequence can be generated for an animal. This means that equine scientists and veterinarians could soon be able to fully and routinely sequence and analyze the genomes of horses in research and clinical settings.
According to Kalbfleisch and MacLeod, the possibilities now exist that in the coming years, we will likely be able to predict simple and complex phenotypes, or physical characteristics that may take years to express themselves, while adjusting management and other environmental variables to minimize or even avoid the impact of disease predispositions. Some of the less complex gene-phenotype disease relationships are already well understood. In other words, precision medicine is upon us.
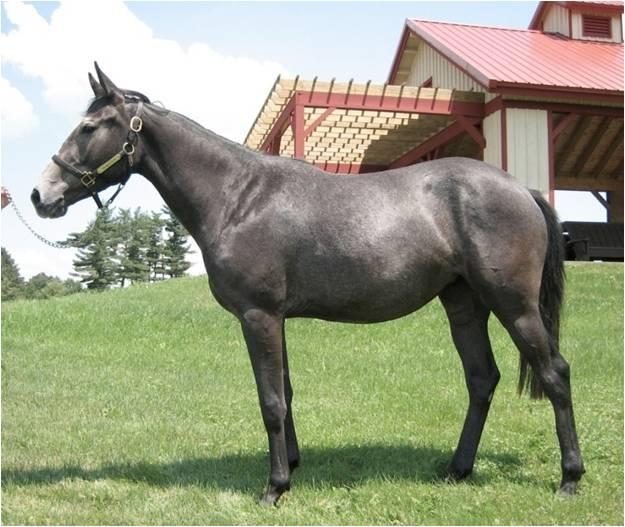

What is big data?
The amount of information created through these studies and by the creation of a genome is enormous. This “big data” and finding patterns in that data is where Kalbfleisch comes in.
He explains the progression of in-formation and the ability to analyze that information.
“Since the late 1990s, the way research is done has been turned a bit on its head,” he said. “We used to ask very specific questions when we would perform measurements to study the molecular biology of living systems. With the limited information we had with respect to the genome, it was difficult to design assays (analyses done to determine the genomic composition of something) that were both sensitive and specific enough to provide the answers needed, but scientists persevered.”
Kalbfleisch said that the technology that allowed researchers to study tens of thousands of genes at once and the data necessary to design the research questions on those data was coming online during that time.
“Now instead of generating a handful of data points, we were generating hundreds of thousands of data points. And then we weren’t simply asking questions about how one gene might behave, but how it and a network of other genes might cooperate in a pathway that drives biological function,” he said. “All of a sudden our notebooks were insufficient, and we needed big disk drives for storage, and big computers to analyze all of this data. About 10 years later, things got more interesting still when we could sequence genomes and transcriptomes in a completely unbiased fashion. At this point, we began generating billions of datapoints to pose even the most routine of research questions.
“Any sensible person would ask, ‘When specific questions worked so well for so long, why spend time and money to generate and deal with all of this data, most of which you won’t need?’ It turns out that it is often less expensive, nearly always faster and always more accurate to generate all of the data than to try to query specific bits of it in the context of a research study,” he said. “And although it is true that much of the data is irrelevant to what is ultimately found to be the answer to the research question at hand, going into the project, we usually have no idea what subset of the data the important bits will be. We know a lot, but what we don’t know dwarfs what we do. And, as important, all of these data are still biologically information rich, and are an incredible resource when made available to the research community through public repositories.”
When asked just how big a change “big data” has brought, Kalbfleisch explained, “I’ve heard it said that the entire recorded history of mankind, up through the year 2000, would have fit into about a terabyte of storage. Now, just about any research project in genetics or genomics will generate at least that much, if not a lot more. My research program alone has 300 terabytes of storage to manage the data my collaborators and I generate. Although it might sound daunting, what has emerged over the last decade in terms of computing power, storage capacity and sophisticated algorithms capable of analyzing all of this data has exceeded our demands. As a result, given a well-defined research question with high quality samples, it is a relatively straightforward process to winnow from these vast quantities of data the information we need to understand the molecular biology of these magnificent animals.”
What are some specific areas of promise using this data for equine medicine?
There are many applications for matching the genetic information found in all this data with the specific medical treatment needed for a single horse. Kalbfleisch outlined a few examples, “Some relatively low hanging fruit one could imagine are more precise targeting of antibiotics. If, for example, you could know for certain what strain of bacteria was making your horse sick and you could avoid using an antibiotic to which that strain of bacteria was known to be resistant, that would be a very good thing. If you could identify a gene or genes that were transcribed at high levels when a horse was experiencing an unhealthy level of stress on bones or joints, you could opt to rest it until it was healthy. If you know of a particular bad trait that runs in a pedigree, you could look at the genomes of affected and unaffected individuals in the pedigree and identify the genetic component responsible for the trait, and in turn make herd management decisions based on whether or not a horse had the genotype for the trait.”
It’s a statistical tool that Kalbfleisch said is a bit like playing a game of “one of these things is not like the other” to determine what traits a population has in common and the traits that might be missing or rare in another population.
Ensuring genetic diversity in the data
One thing Kalbfleisch has been working to improve in the area of genome resources is to add in more genetic diversity. Consider that the Equine Reference Genome is based on a Thoroughbred mare from Cornell named Twilight.
According to Kalbfleisch, the reference works very well for Thoroughbreds and other breeds close to the Thoroughbred but has deficiencies when it comes to other breeds of horses that are more distant.
“The fact that these other breeds aren’t well served by the Twilight reference is a serious problem since many of these differences are likely the very genetic components that distinguish these breeds from the Thoroughbred and, as such, are essential to understand if you hope to do genetic research on them,” he said. “The single reference for the horse is analogous the Model T in cars. And much as with the automobile industry, in the near future, we will begin to see a proliferation of new and varied genomes such that ultimately each major breed has its own. It will be difficult for the research community to manage all of this, but as technologies and our computing infrastructures become more powerful, it is certain that these new resources will make it possible for us to better understand the genetic basis of health in horses.”
He and colleagues are now working to collect samples to catalog genetic diversity across North American Thoroughbreds.
“Genetic diversity is important in any species or breed, and very little is known with respect to the population structure of this wonderful breed. We are working to collect samples from as many animals as possible and will begin by fully sequencing the genomes of as many as 100 of them. As resources become available, we will work to analyze as many of the samples as possible so we have an understanding of what the population structure is now so that we can monitor how genetic diversity changes over time,” he said.
Pandora’s box?
When it comes to genetic studies, there’s also sometimes a concern by horse owners and breeders that DNA tests will reveal that a bloodline or particular valued horse has a bad genetic variant (one of two or more versions of a gene). If so, does that mean they shouldn’t breed that animal?
Actually, genetic testing may provide more opportunities than problems for breeders. First, when a stallion is suspected of harboring a deleterious genetic trait, breeders will shun that stallion (unless his offspring are great runners!). However, genetic testing can exonerate the reputation of a suspect stallion. Second, if a stallion is found to carry a genetic variant that is undesirable, breeders can use genetic tests to design matings and produce offspring in a manner to avoid the deleterious trait.
This is the way that Arabian horse breeders approached the discovery of the recessive deleterious variant causing the disease severe combined immunodeficiency (SCID) in Arabian horses. Once a test was invented, they could have culled all carriers. However, the carriers presumably had a lot of good genetic variants. With the use of the test, it was possible to avoid matings that produced SCID and even purchase offspring from SCID carriers that could be certified as free of SCID.
The role of genetics research and genetic testing is to provide breeders with useful information that allows them to make informed decisions about breeding. Genomic research can turn genetics from a black box into a useful tool.
Kalbfleisch also reiterated that researchers work hard to maintain confidentiality of the specific horse genetic information they work with because they do recognize the concerns horse owners and breeders have.
Partnerships with the industry are important
On Kalbfleisch’s wish list for more data? He said it would be to engage more productively with producers and with veterinarians in order to build a conduit for research to help with the problems people have. Having access to samples and really good phenotype information is key.
Excitement about what the future holds
When asked what excites him about his research, Kalbfleisch was quick to respond. “We know the genome of the horse and where the genes are and much about what they do. The cell doesn’t know anything about any of that. It just functions. It reacts according to the physical and chemical laws that govern everything else. What I’d like to do is to move all of this to where we’re no longer applying these biological or biochemical rules, but to where we’re just using more simple physics and chemistry. And just see how a cell is going to behave when it has a genome that looks like your horse’s,” he said.
“It’s just the opportunity to fill in all of those blanks, all this stuff that we don’t know. Maybe not in 10 years, but in my lifetime. That’s exciting,” he said.
Holly Wiemers, MA, APR, is the communications and managing director for UK Ag Equine Programs.
